I guess you have tunneled in on tuning too many non-useful hyper parameters, because an easy to use grid-search functionality allowed you to do so.
Notice all your explained variances only differ on the fourth digit. You have found, what appears to be a negligible better model settings. But even that you cannot be sure off because:
- the RF model is non-deterministic and performance will vary slightly
- a CV only estimates future model performance with a limited precision
- nfold CV is not perfect reproducible and should be repeated to increase precision
- Grid tuning should be performed with nested CV, but that is not your problem here I think.
Only "grid-tune" max_features. It has only 6 possoble values. You can run each 5 times and plot it. Check if some setting is repetitively better, probably you find anything from 2-4 perform fine. Max_depth is by default unlimited and that is optimal as long data is not very noisy. You set it to 25, which in practice is unlimited because already $2^{15}$=32000 and you "only" have 26000 samples. Changing these other hyper parameter will only give you shorter training times(useful) and/or more robust models. Thumb-rule: as explained variance is way above 50%, you do not need to make your model more robust by limiting depth of trees (max_depth, min_samples_split) to e.g. 3. Max_depth 15 is quite deep, and probably plenty deep enough, just as 2000 are trees enough. So raising and lowering number of trees and depth within the quite fine range does not change anything, and it will be really hard and non-rewarding to find the true best setting.
So you have performed a grid search and learned that RF will have the same performance in the parameter space you have tested.
If you obtain a testset from a different source you should expect a drop in performance. Your CV only estimate the model performance, if the future test set was drawn from the exactly same population.
With 1400 tests, the sample error alone could swing the measured performance +/- 0.03, I guess.
If your swapped e.g. to boosting algorithms grid-tuning of multiple parameters would be a more rewarding tool.
To improve your model maybe you can refine your features. Look to variable importance, to see what features work well. Could you maybe derive new features with an even higher variable importance? Since your explained variance is quite high(low noise), you may benefit from swapping to xgboost. You may also spend time wondering if this chase of a better model performance of some target by some metric (explained variance) is useful specifically for your purpose. Maybe you don't need the model being that accurate when predicting large values, so you log transpose your target e.g. Maybe you only want to rank your predictions so explained variance could be replace with Spearman rank coefficient.
happy modelling:)
Stacking ensembles are usually heterogeneous ensembles that use learners of different types. In order for ensemble methods to be more accurate than any of its individual members the base learners have to be as accurate as possible and as diverse as possible. Diversity can be achieved by using different learners, sub-sampling the data and/or the features, or using learners with different parameter settings. It is possible that one of the base learners is a random forest in which case there will be sub-sampling of data and features. A potential danger in using a sub-set of features for each base model is that the accuracy will degrade in favor of greater diversity. This may be dataset specific and may or may not improve the accuracy of the overall ensemble. It seems that cross-validation will give an experimental answer to this question.
Best Answer
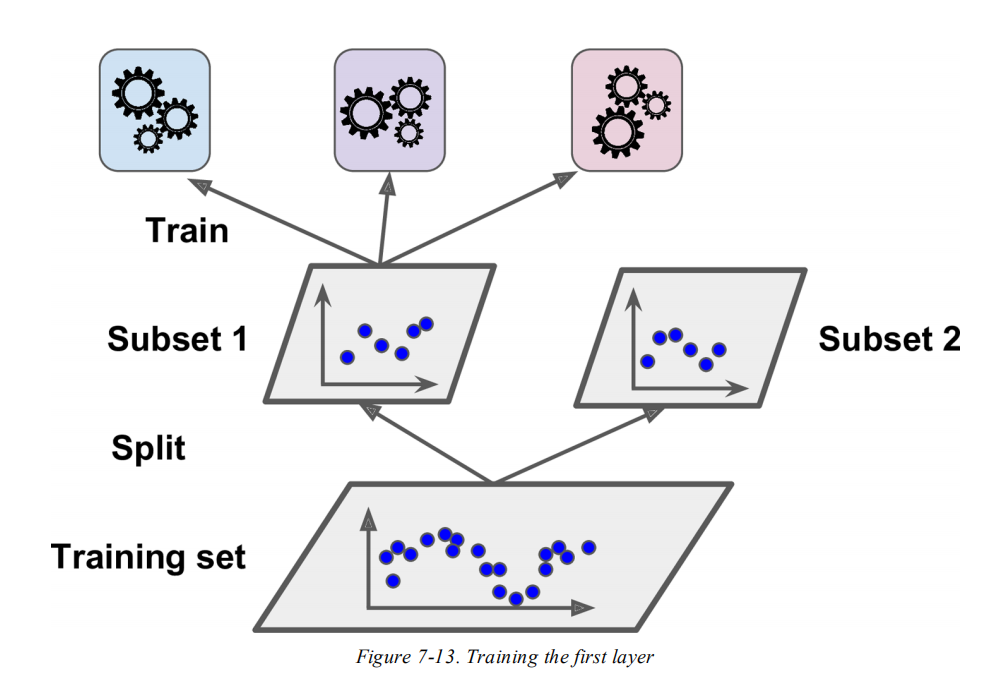
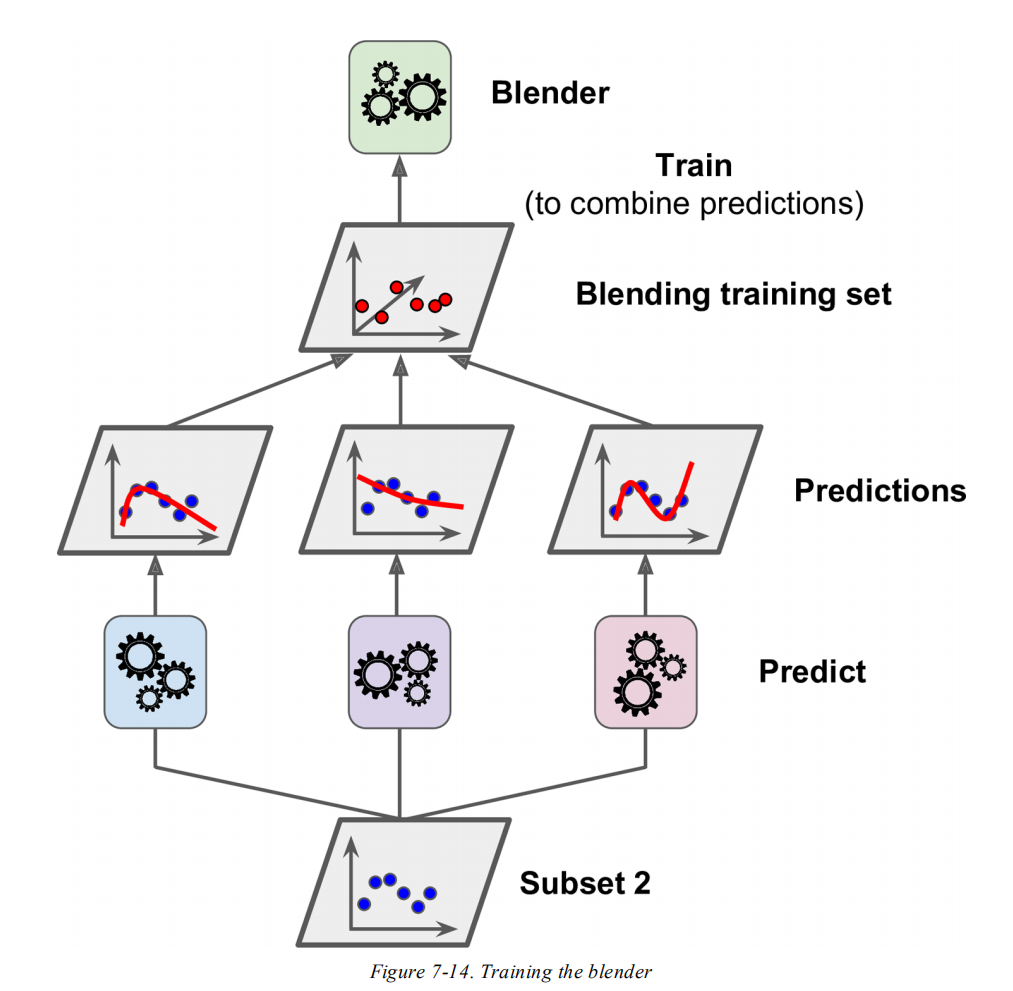
For an exact answer to the why question you would need to ask the scikit-learn developers about their motivation, we could only guess. In general, stacking means training several models on the data and having a meta-model that is trained on the predictions. Aurélien Géron mentions that it is a "common approach is to use a hold-out set", but it is not the only way. Another possible approach is to use cross-validation, in fact, this is what scikit-learn does.
Why did they decide to use cross-validation rather than two separate sets? Cross-validation re-uses the data, so it is better suitable for smaller datasets. For using two sets, you would need a large dataset so that each of the subsets would be big enough to train a model. It seems to be a safer default for general use software, while as noticed by Aurélien Géron, the other approach can be easily implemented if you want to use it.